It seems as if I’ve been writing up nanomedicine research a lot lately, so I would have avoided this piece. However, since I do try to cover Canadian nanotechnology regardless of the topic and this work features researchers from l’Université de Montréal (Québec, Canada), here’s one of the latest innovations in the field of nanomedicine. (I have some additional comments about the nano scene in Canada and one major issue concerning nanomedicine at the end of this posting.) From a May 8, 2017 news item on ScienceDaily,
An international team of researchers from the University of Rome Tor Vergata and the University of Montreal has reported, in a paper published this week in Nature Communications, the design and synthesis of a nanoscale molecular slingshot made of DNA that is 20,000 times smaller than a human hair. This molecular slingshot could “shoot” and deliver drugs at precise locations in the human body once triggered by specific disease markers.
A May 8, 2017 University of Montreal news release (also on EurekAlert), which originated the news item, delves further into the research (Note: A link has been removed),
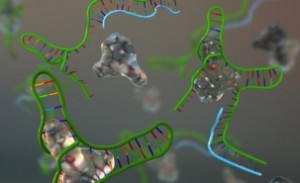
The molecular slingshot is only a few nanometres long and is composed of a synthetic DNA strand that can load a drug and then effectively act as the rubber band of the slingshot. The two ends of this DNA “rubber band” contain two anchoring moieties that can specifically stick to a target antibody, a Y-shaped protein expressed by the body in response to different pathogens such as bacteria and viruses. When the anchoring moieties of the slingshot recognize and bind to the arms of the target antibody the DNA “rubber band” is stretched and the loaded drug is released.
“One impressive feature about this molecular slingshot,” says Francesco Ricci, Associate Professor of Chemistry at the University of Rome Tor Vergata, “is that it can only be triggered by the specific antibody recognizing the anchoring tags of the DNA ‘rubber band’. By simply changing these tags, one can thus program the slingshot to release a drug in response to a variety of specific antibodies. Since different antibodies are markers of different diseases, this could become a very specific weapon in the clinician’s hands.”
“Another great property of our slingshot,” adds Alexis Vallée-Bélisle, Assistant Professor in the Department of Chemistry at the University of Montreal, “is its high versatility. For example, until now we have demonstrated the working principle of the slingshot using three different trigger antibodies, including an HIV antibody, and employing nucleic acids as model drugs. But thanks to the high programmability of DNA chemistry, one can now design the DNA slingshot to ‘shoot’ a wide range of threrapeutic molecules.”
“Designing this molecular slingshot was a great challenge,” says Simona Ranallo, a postdoctoral researcher in Ricci’s team and principal author of the new study. “It required a long series of experiments to find the optimal design, which keeps the drug loaded in ‘rubber band’ in the absence of the antibody, without affecting too much its shooting efficiency once the antibody triggers the slingshot.”
The group of researchers is now eager to adapt the slingshot for the delivery of clinically relevant drugs, and to demonstrate its clinical efficiency. [emphasis mine] “We envision that similar molecular slingshots may be used in the near future to deliver drugs to specific locations in the body. This would drastically improve the efficiency of drugs as well as decrease their toxic secondary effects,” concludes Ricci.
Here’s a link to and a citation for the paper,
Antibody-powered nucleic acid release using a DNA-based nanomachine by Simona Ranallo, Carl Prévost-Tremblay, Andrea Idili, Alexis Vallée-Bélisle, & Francesco Ricci. Nature Communications 8, Article number: 15150 (2017) doi:10.1038/ncomms15150 Published online: 08 May 2017
This is an open access paper.
A couple of comments
The Canadian nanotechnology scene is pretty much centered in Alberta and Québec. The two provinces have invested a fair amount of money in their efforts. Despite the fact that the province of Alberta also hosts the federal government’s National Institute of Nanotechnology, it seems that the province of Québec is the one making the most progress in its various ‘nano’ fields of endeavour. Another province that should be mentioned with regard to its ‘nano’ efforts is Ontario. As far as I can tell, nanotechnology there doesn’t enjoy the same level of provincial funding support as the other two but there is some important work coming out of Ontario.
My other comment has to do with nanomedicine. While it is an exciting field, there is a tendency toward a certain hyperbole. For anyone who got excited about the ‘slingshot’, don’t forget this hasn’t been tested on any conditions close to the conditions found in a human body nor have they even used, “... clinically relevant drugs, … .” It’s also useful to know that less than 1% of the drugs used in nanoparticle-delivery systems make their way to the affected site (from an April 27, 2016 posting about research investigating the effectiveness of nanoparticle-based drug delivery systems). By the way, it was a researcher at the University of Toronto (Ontario, Canada) who first noted this phenomenon after a meta-analysis of the research,
…
More generally, the authors argue that, in order to increase nanoparticle delivery efficiency, a systematic and coordinated long-term strategy is necessary. To build a strong foundation for the field of cancer nanomedicine, researchers will need to understand a lot more about the interactions between nanoparticles and the body’s various organs than they do today. …
It’s not clear from the news release, the paper, or the May 8, 2017 article by Sherry Noik for the Canadian Broadcasting Corporation’s News Online website, how this proposed solution would be administered but presumably the same factors which affect other nano-based drug deliveries could affect this new one,
Scientists have for many years been working on improving therapies like chemo and radiation on that score, but most efforts have focused on modifying the chemistry rather than altering the delivery of the drug.
“It’s all about tuning the concentration of the drug optimally in the body: high concentration where you want it to be active, and low concentration where you don’t want to affect other healthy parts,” says Prof. Alexis Vallée-Bélisle of the University of Montreal, co-author of the report published this week in Nature Communications.
“If you can increase the concentration of that drug at the specific location, that drug will be more efficient,” he told CBC News in an interview.
‘Like a weapon’
Restricting the movement of the drug also reduces potentially harmful secondary effects on other parts of the body — for instance, the hair loss that can result from toxic cancer treatments, or the loss of so-called good bacteria due to antibiotic use.
The idea of the slingshot is to home in on the target cells at a molecular level.
…
The two ends of the strand anchor themselves to the antibody, stretching the strand taut and catapulting the drug to its target.
“Imagine our slingshot like a weapon, and this weapon is being used by our own antibody,” said Vallée-Bélisle, who heads the Laboratory of Biosensors & Nanomachines at U of M. “We design a specific weapon targeting, for example, HIV. We provide the weapon in the body with the bullet — the drug. If the right solider is there, the soldier can use the weapon and shoot the problem.”
Equally important: if the wrong soldier is present, the weapon won’t be deployed.
So rather than delay treatment for an unidentified infection that could be either viral or bacterial, a patient could receive the medication for both and their body would only use the one it needed.
Getting back to my commentary, how does the drug get to its target? Through the bloodstream? Does it get passed through various organs? How do we increase the amount of medication (in nano-based drug delivery systems) reaching affected areas from less than 1%?
The researchers deserve to be congratulated for this work and given much encouragement and thanks as they grapple with the questions I’ve posed and with all of the questions I don’t know how to ask.