The approach looks promising but there’s a still long way to go before this ‘simpler, gentler’ approach to gene therapy will make its way into any treatments. From an August 30, 2017 news item on Nanowerk,
A new biomedical tool using nanoparticles that deliver transient gene changes to targeted cells could make therapies for a variety of diseases — including cancer, diabetes and HIV — faster and cheaper to develop, and more customizable.
The tool, developed by researchers at Fred Hutchinson Cancer Research Center and tested in preclinical models, is described in a paper published August 30 [2017] in Nature Communications.
This animation demonstrates the approach,
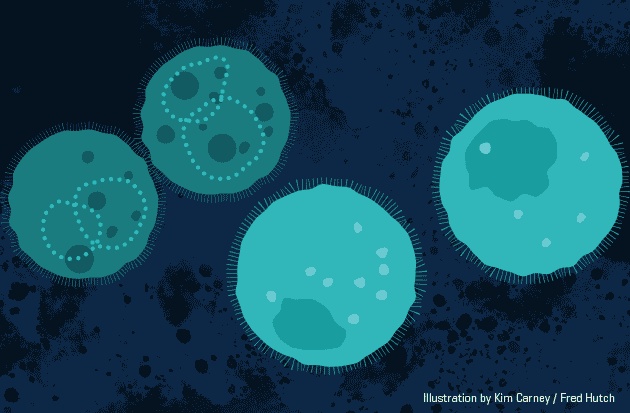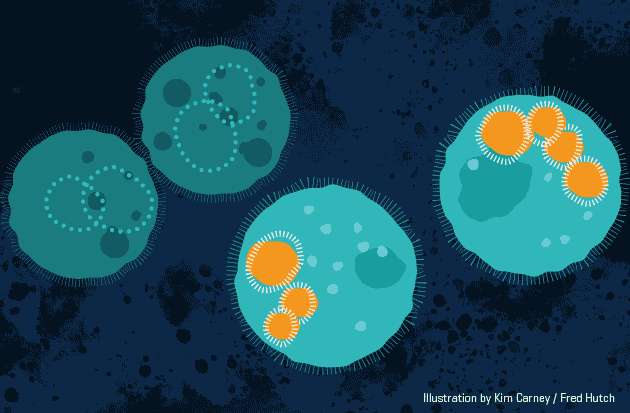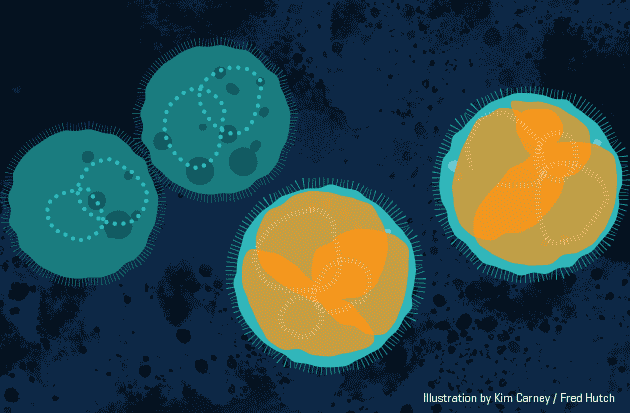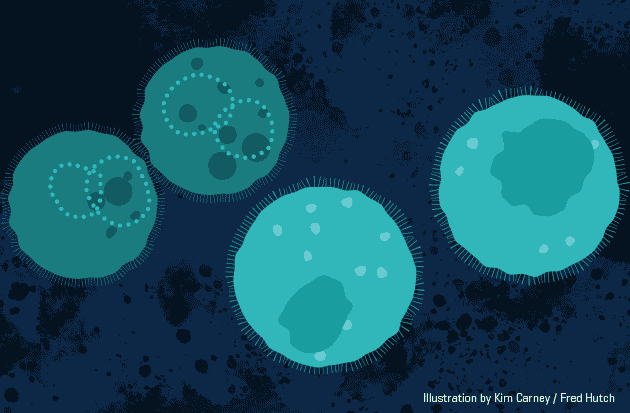
Biodegradable nanoparticles (orange) carry short-lived gene therapy to specific cells (light teal). Animation by Kimberly Carney / Fred Hutch News Service
An August 30, 2017 Fred Hutchinson Cancer Research Center (Fred Hutch) news release (from news release received via email; also on EurekAlert) by Sabrina Richards, which originated the news item, elucidates further (Note: Some links and notes have been removed),
“Our goal is to streamline the manufacture of cell-based therapies,” said lead author DR. MATTHIAS STEPHAN [6], a faculty member in the Fred Hutch Clinical Research Division and an expert in developing biomaterials. “In this study, we created a product where you just add it to cultured cells and that’s it — no additional manufacturing steps.”
Stephan and his colleagues developed a nanoparticle delivery system to extend the therapeutic potential of messenger RNA, which delivers molecular instructions from DNA to cells in the body, directing them to make proteins to prevent or fight disease.
The researchers’ approach was designed to zero in on specific cell types — T cells of the immune system and blood stem cells — and deliver mRNA directly to the cells, triggering short-term gene expression. It’s called “hit-and-run” genetic programming because the transient effect of mRNA does not change the DNA, but it is enough to make a permanent impact on the cells’ therapeutic potential.
Stephan and colleagues used three examples in the Nature Communications paper to demonstrate their technology:
* Nanoparticles carried a gene-editing tool to T cells of the immune system that snipped out their natural T-cell receptors, and then was paired with genes encoding a “chimeric antigen receptor” or CAR, a synthetic molecule designed to attack cancer.
* Targeted to blood stem cells, nanoparticles were equipped with mRNA that enabled the stem cells to multiply and replace blood cancer cells with healthy cells when used in bone marrow transplants.
* Nanoparticles targeted to CAR-T cells and containing foxo1 mRNA, which signals the anti-cancer T cells to develop into a type of “memory” cell that is more aggressive and destroys tumor cells more effectively and maintains anti-tumor activity longer.Other attempts to engineer mRNA into disease-fighting cells have been tricky. The large messenger molecule degrades quickly before it can have an effect, and the body’s immune system recognizes it as foreign — not coming from DNA in the nucleus of the cell — and destroys it.
Stephan and his Fred Hutch collaborators devised a workaround to those hurdles.
“We developed a nanocarrier that binds and condenses synthetic mRNA and protects it from degradation,” Stephan said. The researchers surrounded the nanoparticle with a negatively charged envelope with a targeting ligand attached to the surface so that the particle selectively homes in and binds to a particular cell type.
The cells swallow up the tiny carrier, which can be loaded with different types of manmade mRNA. “If you know from the scientific literature that a signaling pathway works in synergy, you could co-deliver mRNA in a single nanoparticle,” Stephan said. “Every cell that takes up the nanoparticle can express both.”
The approach involves mixing the freeze-dried nanoparticles with water and a sample of cells. Within four hours, cells start showing signs that the editing has taken effect. Boosters can be given if needed. Made from a dissolving biomaterial, the nanoparticles are removed from the body like other cell waste.
“Just add water to our freeze-dried product,” Stephan said. Since it’s built on existing technologies and doesn’t require knowledge of nanotechnology, he intends for it to be an off-the-shelf way for cell-therapy engineers to develop new approaches to treating a variety of diseases.
The approach could replace labor-intensive electroporation, a multistep cell-manufacturing technique that requires specialized equipment and clean rooms. All the handling ends up destroying many of the cells, which limits the amount that can be used in treatments for patients.
Gentler to cells, the nanoparticle system developed by the Fred Hutch team showed that up to 60 times more cells survive the process compared with electroporation. This is a critical feature for ensuring enough cells are viable when transferred to patients.
“You can imagine taking the nanoparticles, injecting them into a patient and then you don’t have to culture cells at all anymore,” he said.
Stephan has tested the technology is cultured cells in the lab, and it’s not yet available as a treatment. Stephan is looking for commercial partners to move the technology toward additional applications and into clinical trials where it could be developed into a therapy.
Here’s a link to and a citation for the paper,
Hit-and-run programming of therapeutic cytoreagents using mRNA nanocarriers by H. F. Moffett, M. E. Coon, S. Radtke, S. B. Stephan, L. McKnight, A. Lambert, B. L. Stoddard, H. P. Kiem, & M. T. Stephan. Nature Communications 8, Article number: 389 (2017) doi:10.1038/s41467-017-00505-8 Published online: 30 August 2017
This paper is open access.