Usually, there’s a rough chronological order to how I introduce the research, but this time I’m looking at the term used to describe it, following up with the various news releases and commentaries about the research, and finishing with a Canadian perspective.
After writing this post (but before it was published), the Weizmann Institute of Science (Israel) made their September 6, 2023 announcement and things changed a bit. That’s in Part two.
Say what you really mean (a terminology issue)
First, it might be useful to investigate the term, ‘synthetic human embryos’ as Julian Hitchcock does in his June 29, 2023 article on Bristows website (h/t Mondaq’s July 5, 2023 news item), Note: Links have been removed,
“Synthetic Embryos” are neither Synthetic nor Embryos. So why are editors giving that name to stem cell-based models of human development?
One of the less convincing aspects of the last fortnight’s flurry of announcements about advances in simulating early human development (see here) concerned their name. Headlines galore (in newspapers and scientific journals) referred to “synthetic embryos“.
But embryo models, however impressive, are not embryos. To claim that the fundamental stages of embryo development that we learnt at school – fertilisation, cleavage and compaction – could now be bypassed to achieve the same result would be wrong. Nor are these objects “synthesised”: indeed, their interest to us lies in the ways in which they organise themselves. The researchers merely place the stem cells in a matrix in appropriate conditions, then stand back and watch them do it. Scientists were therefore unhappy about this use of the term in news media, and relieved when the International Society for Stem Cell Research (ISSCR) stepped in with a press release:
“Unlike some recent media reports describing this research, the ISSCR advises against using the term “synthetic embryo” to describe embryo models, because it is inaccurate and can create confusion. Integrated embryo models are neither synthetic nor embryos. While these models can replicate aspects of the early-stage development of human embryos, they cannot and will not develop to the equivalent of postnatal stage humans. Further, the ISSCR Guidelines prohibit the transfer of any embryo model to the uterus of a human or an animal.”
Although this was the ISSCR’s first attempt to put that position to the public, it had already made that recommendation to the research community two years previously. Its 2021 Guidelines for Stem Cell Research and Clinical Translation had recommended researchers to “promote accurate, current, balanced, and responsive public representations of stem cell research”. In particular:
“While organoids, chimeras, embryo models, and other stem cell-based models are useful research tools offering possibilities for further scientific progress, limitations on the current state of scientific knowledge and regulatory constraints must be clearly explained in any communications with the public or media. Suggestions that any of the current in vitro models can recapitulate an intact embryo, human sentience or integrated brain function are unfounded overstatements that should be avoided and contradicted with more precise characterizations of current understanding.”
…
Here’s a little bit about Hitchcock from his Bristows profile page,
- Diploma Medical School, University of Birmingham (1975-78)
- LLB, University of Wolverhampton
- Diploma in Intellectual Property Law & Practice, University of Bristol
- Qualified 1998
Following an education in medicine at the University of Birmingham and a career as a BBC science producer, Julian has focused on the law and regulation of life science technologies since 1997, practising in England and Australia. He joined Bristows with Alex Denoon in 2018.
…
Hitchcock’s June 29, 2023 article comments on why this term is being used,
…
I have a lot of sympathy with the position of the science writers and editors incurring the scientists’ ire. First, why should journalists have known of the ISSCR’s recommendations on the use of the term “synthetic embryo”? A journalist who found Recommendation 4.1 of the ISSCR Guidelines would probably not have found them specific enough to address the point, and the academic introduction containing the missing detail is hard to find. …
My second reason for being sympathetic to the use of the terrible term is that no suitable alternative has been provided, other than in the Stem Cell Reports paper, which recommends the umbrella terms “embryo models” or “stem cell based embryo models”. …
When asked why she had used the term “synthetic embryo”, the journalist I contacted remarked that, “We’re still working out the right language and it’s something we’re discussing and will no doubt evolve along with the science”.
It is absolutely in the public’s interest (and in the interest of science), that scientific research is explained in terms that the public understands. There is, therefore, a need, I think, for the scientific community to supply a name to the media or endure the penalties of misinformation …
In such an intensely competitive field of research, disagreement among researchers, even as to names, is inevitable. In consequence, however, journalists and their audiences are confronted by a slew of terms which may or may not be synonymous or overlapping, with no agreed term [emphasis mine] for the overall class of stem cell based embryo models. We cannot blame them if they make up snappy titles of their own [emphasis mine]. …
The announcement
The earliest date for the announcement at the International Society for Stem Cell Researh meeting that I can find is Hannah Devlin’s June 14, 2023 article in The Guardian newspaper, Note: A link has been removed,
Scientists have created synthetic human embryos using stem cells, in a groundbreaking advance that sidesteps the need for eggs or sperm.
Scientists say these model embryos, which resemble those in the earliest stages of human development, could provide a crucial window on the impact of genetic disorders and the biological causes of recurrent miscarriage.
However, the work also raises serious ethical and legal issues as the lab-grown entities fall outside current legislation in the UK and most other countries.
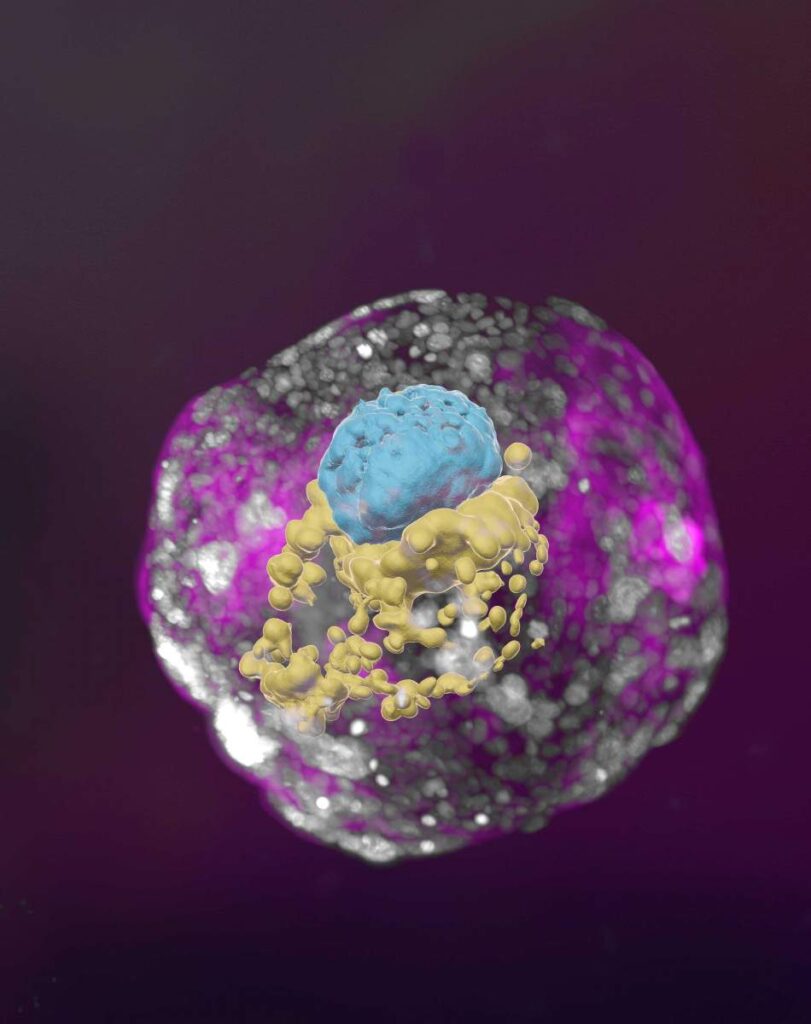
The structures do not have a beating heart or the beginnings of a brain, but include cells that would typically go on to form the placenta, yolk sac and the embryo itself.
Prof Magdalena Żernicka-Goetz, of the University of Cambridge and the California Institute of Technology, described the work in a plenary address on Wednesday [June 14, 2023] at the International Society for Stem Cell Research’s annual meeting in Boston.
…
The (UK) Science Media Centre made expert comments available in a June 14, 2023 posting “expert reaction to Guardian reporting news of creation of synthetic embryos using stem cells.”
Two days later, this June 16, 2023 essay by Kathryn MacKay, Senior Lecturer in Bioethics, University of Sydney (Australia), appeared on The Conversation (h/t June 16, 2023 news item on phys.org), Note: Links have been removed,
Researchers have created synthetic human embryos using stem cells, according to media reports. Remarkably, these embryos have reportedly been created from embryonic stem cells, meaning they do not require sperm and ova.
This development, widely described as a breakthrough that could help scientists learn more about human development and genetic disorders, was revealed this week in Boston at the annual meeting of the International Society for Stem Cell Research.
The research, announced by Professor Magdalena Żernicka-Goetz of the University of Cambridge and the California Institute of Technology, has not yet been published in a peer-reviewed journal. But Żernicka-Goetz told the meeting these human-like embryos had been made by reprogramming human embryonic stem cells.
So what does all this mean for science, and what ethical issues does it present?
MacKay goes on to answer her own questions, from the June 16, 2023 essay, Note: A link has been removed,
…
One of these quandaries arises around whether their creation really gets us away from the use of human embryos.
Robin Lovell-Badge, the head of stem cell biology and developmental genetics at the Francis Crick Institute in London UK, reportedly said that if these human-like embryos can really model human development in the early stages of pregnancy, then we will not have to use human embryos for research.
At the moment, it is unclear if this is the case for two reasons.
First, the embryos were created from human embryonic stem cells, so it seems they do still need human embryos for their creation. Perhaps more light will be shed on this when Żernicka-Goetz’s research is published.
Second, there are questions about the extent to which these human-like embryos really can model human development.
…
Professor Magdalena Żernicka-Goetz’s research is published
Almost two weeks later the research from the Cambridge team (there are other teams and countries also racing; see Part two for the news from Sept. 6, 2023) was published, from a June 27, 2023 news item on ScienceDaily,
Cambridge scientists have created a stem cell-derived model of the human embryo in the lab by reprogramming human stem cells. The breakthrough could help research into genetic disorders and in understanding why and how pregnancies fail.
Published today [Tuesday, June 27, 2023] in the journal Nature, this embryo model is an organised three-dimensional structure derived from pluripotent stem cells that replicate some developmental processes that occur in early human embryos.
Use of such models allows experimental modelling of embryonic development during the second week of pregnancy. They can help researchers gain basic knowledge of the developmental origins of organs and specialised cells such as sperm and eggs, and facilitate understanding of early pregnancy loss.
…
A June 27, 2023 University of Cambridge press release (also on EurekAlert), which originated the news item, provides more detail about the work,
“Our human embryo-like model, created entirely from human stem cells, gives us access to the developing structure at a stage that is normally hidden from us due to the implantation of the tiny embryo into the mother’s womb,” said Professor Magdalena Zernicka-Goetz in the University of Cambridge’s Department of Physiology, Development and Neuroscience, who led the work.
She added: “This exciting development allows us to manipulate genes to understand their developmental roles in a model system. This will let us test the function of specific factors, which is difficult to do in the natural embryo.”
In natural human development, the second week of development is an important time when the embryo implants into the uterus. This is the time when many pregnancies are lost.
The new advance enables scientists to peer into the mysterious ‘black box’ period of human development – usually following implantation of the embryo in the uterus – to observe processes never directly observed before.
Understanding these early developmental processes holds the potential to reveal some of the causes of human birth defects and diseases, and to develop tests for these in pregnant women.
Until now, the processes could only be observed in animal models, using cells from zebrafish and mice, for example.
Legal restrictions in the UK currently prevent the culture of natural human embryos in the lab beyond day 14 of development: this time limit was set to correspond to the stage where the embryo can no longer form a twin. [emphasis mine]
Until now, scientists have only been able to study this period of human development using donated human embryos. This advance could reduce the need for donated human embryos in research.
Zernicka-Goetz says the while these models can mimic aspects of the development of human embryos, they cannot and will not develop to the equivalent of postnatal stage humans.
Over the past decade, Zernicka-Goetz’s group in Cambridge has been studying the earliest stages of pregnancy, in order to understand why some pregnancies fail and some succeed.
In 2021 and then in 2022 her team announced in Developmental Cell, Nature and Cell Stem Cell journals that they had finally created model embryos from mouse stem cells that can develop to form a brain-like structure, a beating heart, and the foundations of all other organs of the body.
The new models derived from human stem cells do not have a brain or beating heart, but they include cells that would typically go on to form the embryo, placenta and yolk sac, and develop to form the precursors of germ cells (that will form sperm and eggs).
Many pregnancies fail at the point when these three types of cells orchestrate implantation into the uterus begin to send mechanical and chemical signals to each other, which tell the embryo how to develop properly.
There are clear regulations governing stem cell-based models of human embryos and all researchers doing embryo modelling work must first be approved by ethics committees. Journals require proof of this ethics review before they accept scientific papers for publication. Zernicka-Goetz’s laboratory holds these approvals.
“It is against the law and FDA regulations to transfer any embryo-like models into a woman for reproductive aims. These are highly manipulated human cells and their attempted reproductive use would be extremely dangerous,” said Dr Insoo Hyun, Director of the Center for Life Sciences and Public Learning at Boston’s Museum of Science and a member of Harvard Medical School’s Center for Bioethics.
Zernicka-Goetz also holds position at the California Institute of Technology and is NOMIS Distinguished Scientist and Scholar Awardee.
The research was funded by the Wellcome Trust and Open Philanthropy.
(There’s more about legal concerns further down in this post.)
Here’s a link to and a citation for the paper,
Pluripotent stem cell-derived model of the post-implantation human embryo by Bailey A. T. Weatherbee, Carlos W. Gantner, Lisa K. Iwamoto-Stohl, Riza M. Daza, Nobuhiko Hamazaki, Jay Shendure & Magdalena Zernicka-Goetz. Nature (2023) DOI: https://doi.org/10.1038/s41586-023-06368-y Published: 27 June 2023
This paper is open access.
Published the same day (June 27, 2023) is a paper (citation and link follow) also focused on studying human embryonic development using stem cells. First, there’s this from the Abstract,
Investigating human development is a substantial scientific challenge due to the technical and ethical limitations of working with embryonic samples. In the face of these difficulties, stem cells have provided an alternative to experimentally model inaccessible stages of human development in vitro …
This time the work is from a US/German team,
Self-patterning of human stem cells into post-implantation lineages by Monique Pedroza, Seher Ipek Gassaloglu, Nicolas Dias, Liangwen Zhong, Tien-Chi Jason Hou, Helene Kretzmer, Zachary D. Smith & Berna Sozen. Nature (2023) DOI: https://doi.org/10.1038/s41586-023-06354-4 Published: 27 June 2023
The paper is open access.
Legal concerns and a Canadian focus
A July 25, 2023 essay by Françoise Baylis and Jocelyn Downie of Dalhousie University (Nova Scotia, Canada) for The Conversation (h/t July 25, 2023 article on phys.org) covers the advantages of doing this work before launching into a discussion of legislation and limits in the UK and, more extensively, in Canada, Note: Links have been removed,
…
This research could increase our understanding of human development and genetic disorders, help us learn how to prevent early miscarriages, lead to improvements in fertility treatment, and — perhaps — eventually allow for reproduction without using sperm and eggs.
Synthetic human embryos — also called embryoid bodies, embryo-like structures or embryo models — mimic the development of “natural human embryos,” those created by fertilization. Synthetic human embryos include the “cells that would typically go on to form the embryo, placenta and yolk sac, and develop to form the precursors of germ cells (that will form sperm and eggs).”
Though research involving natural human embryos is legal in many jurisdictions, it remains controversial. For some people, research involving synthetic human embryos is less controversial because these embryos cannot “develop to the equivalent of postnatal stage humans.” In other words, these embryos are non-viable and cannot result in live births.
…
Now, for a closer look at the legalities in the UK and in Canada, from the July 25, 2023 essay, Note: Links have been removed,
The research presented by Żernicka-Goetz at the ISSCR meeting took place in the United Kingdom. It was conducted in accordance with the Human Fertilization and Embryology Act, 1990, with the approval of the U.K. Stem Cell Bank Steering Committee.
U.K. law limits the research use of human embryos to 14 days of development. An embryo is defined as “a live human embryo where fertilisation is complete, and references to an embryo include an egg in the process of fertilisation.”
Synthetic embryos are not created by fertilization and therefore, by definition, the 14-day limit on human embryo research does not apply to them. This means that synthetic human embryo research beyond 14 days can proceed in the U.K.
The door to the touted potential benefits — and ethical controversies — seems wide open in the U.K.
…
While the law in the U.K. does not apply to synthetic human embryos, the law in Canada clearly does. This is because the legal definition of an embryo in Canada is not limited to embryos created by fertilization [emphasis mine].
The Assisted Human Reproduction Act (the AHR Act) defines an embryo as “a human organism during the first 56 days of its development following fertilization or creation, excluding any time during which its development has been suspended.”
Based on this definition, the AHR Act applies to embryos created by reprogramming human embryonic stem cells — in other words, synthetic human embryos — provided such embryos qualify as human organisms.
A synthetic human embryo is a human organism. It is of the species Homo sapiens, and is thus human. It also qualifies as an organism — a life form — alongside other organisms created by means of fertilization, asexual reproduction, parthenogenesis or cloning.
…
Given that the AHR Act applies to synthetic human embryos, there are legal limits on their creation and use in Canada.
First, human embryos — including synthetic human embryos – can only be created for the purposes of “creating a human being, improving or providing instruction in assisted reproduction procedures.”
Given the state of the science, it follows that synthetic human embryos could legally be created for the purpose of improving assisted reproduction procedures.
Second, “spare” or “excess” human embryos — including synthetic human embryos — originally created for one of the permitted purposes, but no longer wanted for this purpose, can be used for research. This research must be done in accordance with the consent regulations which specify that consent must be for a “specific research project.”
Finally, all research involving human embryos — including synthetic human embryos — is subject to the 14-day rule. The law stipulates that: “No person shall knowingly… maintain an embryo outside the body of a female person after the fourteenth day of its development following fertilization or creation, excluding any time during which its development has been suspended.”
Putting this all together, the creation of synthetic embryos for improving assisted human reproduction procedures is permitted, as is research using “spare” or “excess” synthetic embryos originally created for this purpose — provided there is specific consent and the research does not exceed 14 days.
This means that while synthetic human embryos may be useful for limited research on pre-implantation embryo development, they are not available in Canada for research on post-implantation embryo development beyond 14 days.
The authors close with this comment about the prospects for expanding Canada’s14-day limit, from the July 25, 2023 essay,
… any argument will have to overcome the political reality that the federal government is unlikely to open up the Pandora’s box of amending the AHR Act.
It therefore seems likely that synthetic human embryo research will remain limited in Canada for the foreseeable future.
As mentioned, in September 2023 there was a new development. See: Part two.